
Together with many collaboration partners, Till Andlauer researches the genetics of neurological and psychiatric disorders, with the aim of adding to our understanding of how genetic variants influence risk for disorders, the course and severity of diseases, and the positive and adverse effects of treatments. Most common neurological and psychiatric disorders are highly heterogeneous, which impedes the successful and timely treatment of patients. The application of polygenic risk scores and machine learning methods can help to identify more homogeneous patient subgroups with a shared molecular aetiology. Characterizing these subgroups can support stratified assessments of disorder risk, predictions of individual disease courses, and, eventually, recommendations of targeted therapies.
You may follow my latest research also on Twitter.
Psychiatric genetics
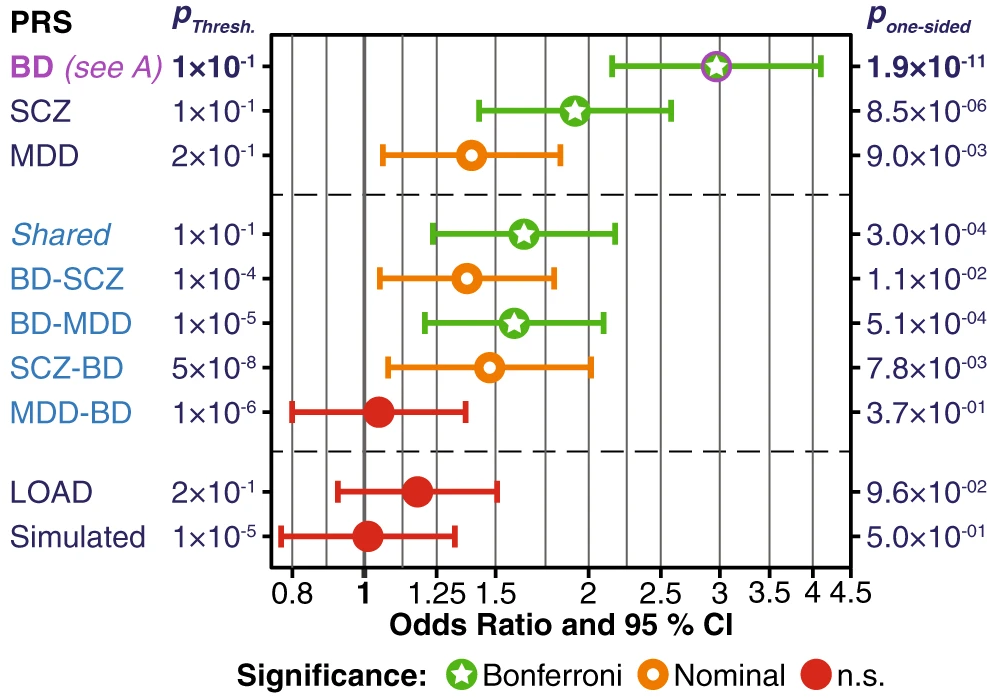
Psychiatric disorders are highly polygenic, and characterizing their risk landscape thus constitutes a challenging task. I am involved in studying the genetics of dyslexia and major depression – see this YouTube video or this review for an introduction to depression genetics. Polygenic risk scores are especially suited to study psychiatric disorders and their subtypes (see our review and my video on the topic).
I have analyzed families with a high prevalence of bipolar disorder and found them to have especially high genetic risk burdens for psychiatric disorders. We also characterized clinical and genetic differences between patients diagnosed with either bipolar type I or II disorder in these families.
Psychiatric symptoms often extend the boundaries of classical diagnostic groups. Thus, we have recently conducted two transdiagnostic analyses: First, a transdiagnostic imaging genetics GWAS and, second, we clustered psychiatric patients and healthy controls using numerous symptom-related variables and then characterized the severity-ranked clusters with polygenic scores.
Genetics of multiple sclerosis
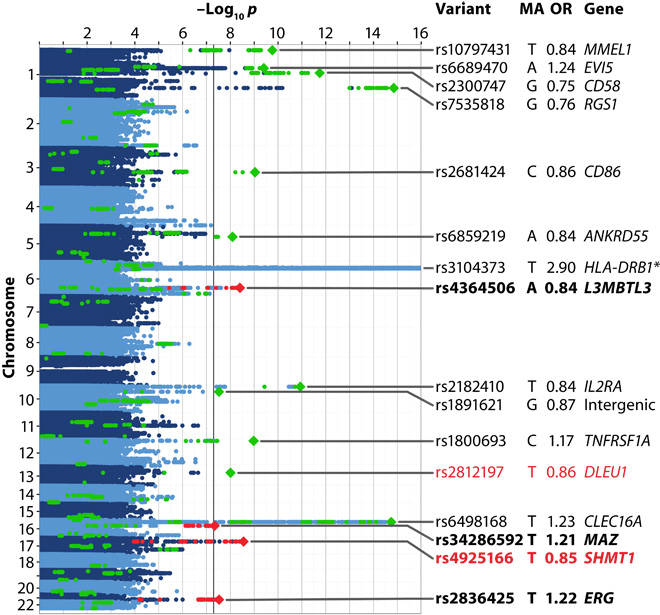
One of my main research interests is the characterization of genetic factors influencing risk for multiple sclerosis or the progression and treatment of the disorder. In 2016, I published a large genome-wide association study on MS susceptibility. See this video for a general introduction to genome-wide association studies. This analysis was followed in 2019 by a more comprehensive study characterizing the genetic map of multiple sclerosis, conducted by the international IMSGC consortium. We have also integrated functional information into GWAS to conduct a novel deep learning-based approach that directly identifies variants functionally relevant for multiple sclerosis. Subsequently, we have analyzed to which degree gene expression in animal models related to multiple sclerosis involves MS risk genes. Recently, we have begun to study the genetics of MS severity, for example, the frequency of relapses.
Pharmacogenomics
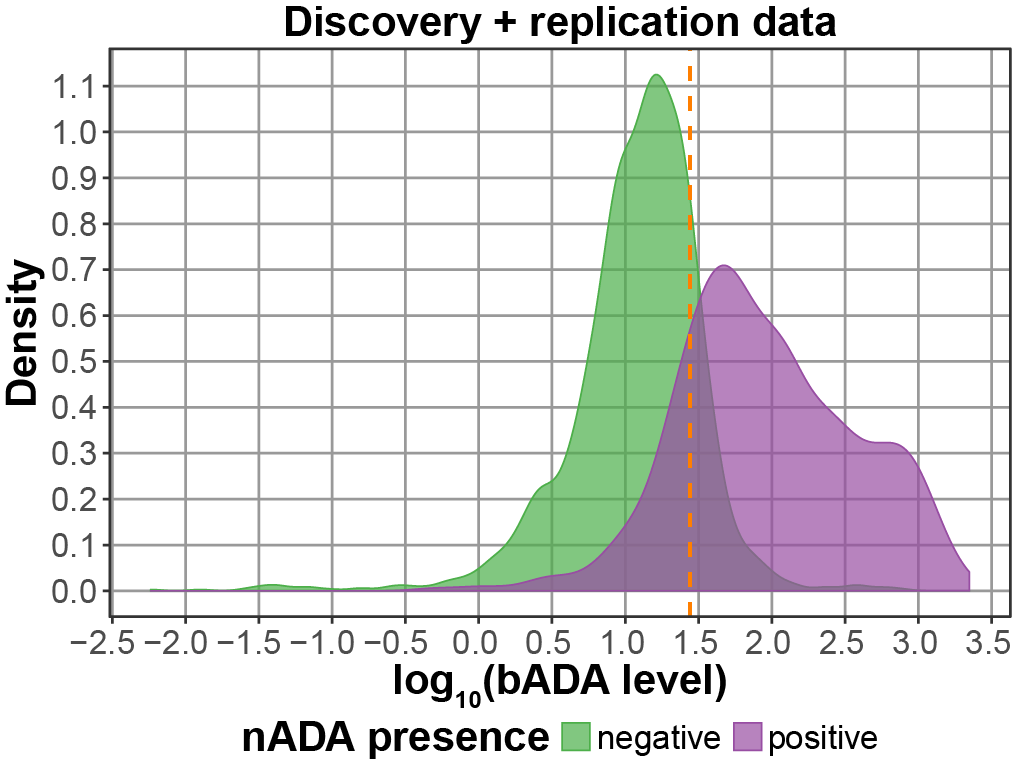
Depending on our individual genetic background, different therapies work more or less effectively in individual patients or cause adverse effects. For any kind of biopharmaceuticals, a patient’s immune system can develop antibodies that neutralize the drug’s function. I have characterized how genetic factors influence the development of such neutralizing anti-drug antibodies against interferon β treatment in MS patients. Following a study published in 2018, I have now conducted a large GWAS for the ABIRISK consortium. See this YouTube video for a presentation of the ABIRSK GWAS project. Clustering methods might help to identify subgroups of patients showing different treatment responses.
Imaging genetics
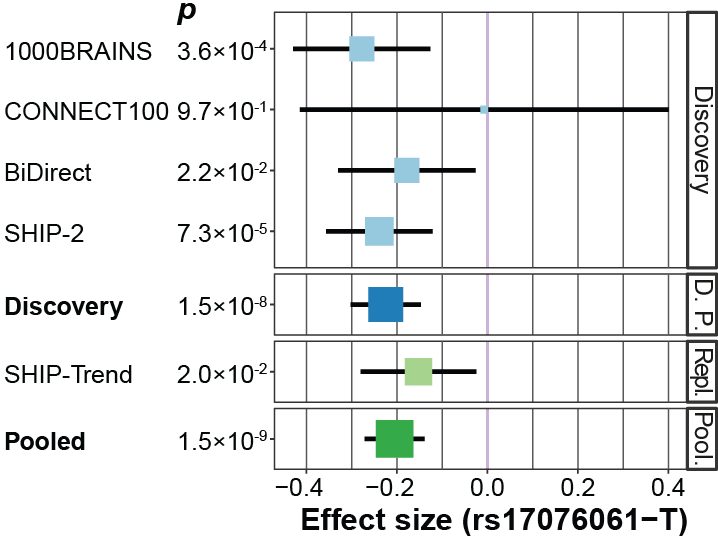
It is well known that genetic variants contribute to the volume, thickness, and surface area of brain regions. I have led a multi-centre analysis of how genetic variants are linked to a neurobiological MRI brain imaging substrate of psychiatric disorders. This transdiagnostic substrate consists of three hubs of the salience network where grey matter reduction is observed in psychiatric patients across the affective-psychotic spectrum. Moreover, genetics likely influence the burden of white matter lesions in MS patients. Together with colleagues from different European consortia, I currently conduct a large GWAS on the white-matter lesion volume in MS patients.
Synaptic plasticity
Many of the risk factors for psychiatric disorders play roles in regulating synaptic plasticity. After describing a postsynaptic gene important for learning and memory, I have examined synaptic diversity in the Drosophila central nervous system. I have also developed analysis pipelines for confocal microscopy, including an ImageJ plugin.
